
ISSN 1004-5759 CN 62-1105/S

Acta Prataculturae Sinica ›› 2026, Vol. 35 ›› Issue (1): 223-240.DOI: 10.11686/cyxb2025063
Miao-miao DOU( ), Xiao-dong JIANG, Hui-qiong SUN, Hong-shen XU, Xi-liang WANG, Bo-hui YANG, Wen-ting CHAI, Shan-shan ZHAO, Chun-lai ZHANG(
), Xiao-dong JIANG, Hui-qiong SUN, Hong-shen XU, Xi-liang WANG, Bo-hui YANG, Wen-ting CHAI, Shan-shan ZHAO, Chun-lai ZHANG( )
)
Received:2025-03-04
Revised:2025-05-06
Online:2026-01-20
Published:2025-11-13
Contact:
Chun-lai ZHANG
Miao-miao DOU, Xiao-dong JIANG, Hui-qiong SUN, Hong-shen XU, Xi-liang WANG, Bo-hui YANG, Wen-ting CHAI, Shan-shan ZHAO, Chun-lai ZHANG. Cloning, expression profiling and DNA variation analysis of the disease-resistance gene CqSGT1 in quinoa (Chenopodium quinoa)[J]. Acta Prataculturae Sinica, 2026, 35(1): 223-240.
基因名称 Gene name | 正向序列 Forward primer (5′→3′) | 反向序列 Reverse primer (5′→3′) | 用途 Application |
|---|---|---|---|
| CqSGT1a | ACGGAAAGGAACTGCGTGTA | GGCTGCTTGTGCATTGGTAG | qRT-PCR, SemiRT-PCR |
| CqSGT1b | TTACCTACGAAAAGGGACCGC | TGCTTGTGCATTGGTAGGCT | qRT-PCR, SemiRT-PCR |
| CqEF1α | GACAAGCGTGTGATCGAGAG | TCGGCCTTAAGTTTGTCGAG | qRT-PCR, SemiRT-PCR |
| TI-CqSGT1a/b | ATGGCGACGGATCTCGAGACCAAG | TTAGTATTCCCATTTCTTCATCTC | 基因克隆Gene clone |
pCAMBIA1300-Cq SGT1a/b | CGC | CGC | 瞬时表达Transient expression |
| M13 | GTTGTAAAACGACGGCCAG | CAGGAAACAGCTATGAC | 基因克隆和瞬时表达Gene clone and transient expression |
Table1 Primers used for gene analysis of PCR
基因名称 Gene name | 正向序列 Forward primer (5′→3′) | 反向序列 Reverse primer (5′→3′) | 用途 Application |
|---|---|---|---|
| CqSGT1a | ACGGAAAGGAACTGCGTGTA | GGCTGCTTGTGCATTGGTAG | qRT-PCR, SemiRT-PCR |
| CqSGT1b | TTACCTACGAAAAGGGACCGC | TGCTTGTGCATTGGTAGGCT | qRT-PCR, SemiRT-PCR |
| CqEF1α | GACAAGCGTGTGATCGAGAG | TCGGCCTTAAGTTTGTCGAG | qRT-PCR, SemiRT-PCR |
| TI-CqSGT1a/b | ATGGCGACGGATCTCGAGACCAAG | TTAGTATTCCCATTTCTTCATCTC | 基因克隆Gene clone |
pCAMBIA1300-Cq SGT1a/b | CGC | CGC | 瞬时表达Transient expression |
| M13 | GTTGTAAAACGACGGCCAG | CAGGAAACAGCTATGAC | 基因克隆和瞬时表达Gene clone and transient expression |
基因名称 Gene name | 基因号 Gene ID | 染色体位置 Chromosome location | 氨基酸长度Length of amino acids (aa) | 分子量 Molecular weight (Da) | 等电点 Isoelectric point (pI) | 不稳定系数Instability coefficient | 脂溶系数 Liposolubility coefficient | 亲水性均值 Grand average of hydropathicity |
|---|---|---|---|---|---|---|---|---|
| CqSGT1a | AUR62021095 | Chr06 | 358 | 39906.09 | 5.18 | 36.21 | 75.03 | -0.515 |
| CqSGT1b | AUR62031443 | Chr07 | 358 | 39890.07 | 5.42 | 34.77 | 73.66 | -0.563 |
Table 2 Basic information of SGT1 family genes in C. quinoa
基因名称 Gene name | 基因号 Gene ID | 染色体位置 Chromosome location | 氨基酸长度Length of amino acids (aa) | 分子量 Molecular weight (Da) | 等电点 Isoelectric point (pI) | 不稳定系数Instability coefficient | 脂溶系数 Liposolubility coefficient | 亲水性均值 Grand average of hydropathicity |
|---|---|---|---|---|---|---|---|---|
| CqSGT1a | AUR62021095 | Chr06 | 358 | 39906.09 | 5.18 | 36.21 | 75.03 | -0.515 |
| CqSGT1b | AUR62031443 | Chr07 | 358 | 39890.07 | 5.42 | 34.77 | 73.66 | -0.563 |
蛋白质名称 Protein name | α-螺旋 α-helix | 延伸链 Extended strand | β-转角 β-turn | 无规则卷曲 Random coil |
|---|---|---|---|---|
| CqSGT1a | 41.62 | 13.41 | 4.19 | 40.78 |
| CqSGT1b | 45.81 | 12.57 | 3.63 | 37.99 |
Table 3 Secondary structures of SGT1 proteins in C. quinoa (%)
蛋白质名称 Protein name | α-螺旋 α-helix | 延伸链 Extended strand | β-转角 β-turn | 无规则卷曲 Random coil |
|---|---|---|---|---|
| CqSGT1a | 41.62 | 13.41 | 4.19 | 40.78 |
| CqSGT1b | 45.81 | 12.57 | 3.63 | 37.99 |
蛋白质 名称 Protein name | 信号肽 Signal peptide | 亚细胞定位 Subcellular localization | 磷酸化位点 Phosphorylation sites | ||
|---|---|---|---|---|---|
| 丝氨酸Serine | 苏氨酸Threonine | 酪氨酸Tyrosine | |||
| CqSGT1a | 无Nothing | 细胞核Nucleus | 16 | 11 | 6 |
| CqSGT1b | 无Nothing | 细胞核Nucleus | 17 | 12 | 6 |
Table 4 Signal peptid, subcellular localization prediction and phosphorylation sites of CqSGT1 proteins in C. quinoa
蛋白质 名称 Protein name | 信号肽 Signal peptide | 亚细胞定位 Subcellular localization | 磷酸化位点 Phosphorylation sites | ||
|---|---|---|---|---|---|
| 丝氨酸Serine | 苏氨酸Threonine | 酪氨酸Tyrosine | |||
| CqSGT1a | 无Nothing | 细胞核Nucleus | 16 | 11 | 6 |
| CqSGT1b | 无Nothing | 细胞核Nucleus | 17 | 12 | 6 |
元件名称 Element name | 核心序列 Core sequence | 元件功能 Element function | 数量Quantity | |
|---|---|---|---|---|
| CqSGT1a | CqSGT1b | |||
| chs-CMA1a | TTACTTAA | 部分光响应元件Part of light responsive element | 1 | - |
| 3-AF1 binding site | TAAGAGAGGAA | 光响应元件Light responsive element | 1 | - |
| HD-Zip 3 | GTAAT(G/C)ATTAC | 蛋白质结合位点Protein binding site | 1 | - |
| ARE | AAACCA | 厌氧诱导必不可少的顺式作用调节元件Cis-acting regulatory element essential for the anaerobic induction | 4 | 1 |
| GT1-motif | GGTTAA | 光响应元件Light responsive element | 2 | - |
| Box 4 | ATTAAT | 参与光响应的部分保守DNA模块Part of conserved DNA module involved in light responsiveness | 3 | 4 |
| G-box | CACGAC | 参与光反应的顺式作用调节元件Cis-acting regulatory element involved in light responsiveness | 1 | - |
| GA-motif | ATAGATAA | 部分光响应元件Part of light responsive element | 1 | - |
| TCT-motif | TCTTAC | 部分光响应元件Part of light responsive element | 1 | - |
| P-box | CCTTTTG | 赤霉素响应元件Gibberellin-responsive element | 2 | 2 |
| Gap-box | CAAATGAA(A/G)A | 光响应元件Light responsive element | 1 | - |
| TCA-element | CCATCTTTTT | 参与水杨酸反应的顺式作用调节件Cis-acting element involved in salicylic acid responsiveness | 1 | - |
| ATCT-motif | AATCTAATCC | 参与光响应的部分保守DNA模块Part of conserved DNA module involved in light responsiveness | 1 | - |
| TATA-box | AATCTAATCC | 转录起始-30核心启动子元件Core promoter element around-30 of transcription start | 64 | 53 |
| MBSI | AAAAAAC(G/C)GTTA | MYB结合位点参与类黄酮生物合成基因的调控元件MYB-binding site as a regulatory element involved in the control of flavonoid biosynthetic genes | - | 1 |
| CAAT-box | CAAAT | 启动子和增强子区域调控元件Promoter and enhancer region regulatory elements | - | 12 |
| TCT-motif | TCTTAC | 部分光响应元件Part of light responsive element | - | 2 |
| O2-site | GTTGACGTGA | 参与玉米醇溶蛋白代谢调控的顺式调节元件Cis-acting regulatory element involved in zein metabolism regulation | - | 1 |
| MRE | AACCTAA | MYB结合位点(参与光响应)MYB binding site involved in light responsiveness | - | 1 |
| Circadian | AACCTAA | 参与昼夜节律控制的顺式作用调节元件Cis-acting regulatory element involved in circadian control | - | 1 |
Table 5 Cis-acting elements of SGT1 genes in C. quinoa
元件名称 Element name | 核心序列 Core sequence | 元件功能 Element function | 数量Quantity | |
|---|---|---|---|---|
| CqSGT1a | CqSGT1b | |||
| chs-CMA1a | TTACTTAA | 部分光响应元件Part of light responsive element | 1 | - |
| 3-AF1 binding site | TAAGAGAGGAA | 光响应元件Light responsive element | 1 | - |
| HD-Zip 3 | GTAAT(G/C)ATTAC | 蛋白质结合位点Protein binding site | 1 | - |
| ARE | AAACCA | 厌氧诱导必不可少的顺式作用调节元件Cis-acting regulatory element essential for the anaerobic induction | 4 | 1 |
| GT1-motif | GGTTAA | 光响应元件Light responsive element | 2 | - |
| Box 4 | ATTAAT | 参与光响应的部分保守DNA模块Part of conserved DNA module involved in light responsiveness | 3 | 4 |
| G-box | CACGAC | 参与光反应的顺式作用调节元件Cis-acting regulatory element involved in light responsiveness | 1 | - |
| GA-motif | ATAGATAA | 部分光响应元件Part of light responsive element | 1 | - |
| TCT-motif | TCTTAC | 部分光响应元件Part of light responsive element | 1 | - |
| P-box | CCTTTTG | 赤霉素响应元件Gibberellin-responsive element | 2 | 2 |
| Gap-box | CAAATGAA(A/G)A | 光响应元件Light responsive element | 1 | - |
| TCA-element | CCATCTTTTT | 参与水杨酸反应的顺式作用调节件Cis-acting element involved in salicylic acid responsiveness | 1 | - |
| ATCT-motif | AATCTAATCC | 参与光响应的部分保守DNA模块Part of conserved DNA module involved in light responsiveness | 1 | - |
| TATA-box | AATCTAATCC | 转录起始-30核心启动子元件Core promoter element around-30 of transcription start | 64 | 53 |
| MBSI | AAAAAAC(G/C)GTTA | MYB结合位点参与类黄酮生物合成基因的调控元件MYB-binding site as a regulatory element involved in the control of flavonoid biosynthetic genes | - | 1 |
| CAAT-box | CAAAT | 启动子和增强子区域调控元件Promoter and enhancer region regulatory elements | - | 12 |
| TCT-motif | TCTTAC | 部分光响应元件Part of light responsive element | - | 2 |
| O2-site | GTTGACGTGA | 参与玉米醇溶蛋白代谢调控的顺式调节元件Cis-acting regulatory element involved in zein metabolism regulation | - | 1 |
| MRE | AACCTAA | MYB结合位点(参与光响应)MYB binding site involved in light responsiveness | - | 1 |
| Circadian | AACCTAA | 参与昼夜节律控制的顺式作用调节元件Cis-acting regulatory element involved in circadian control | - | 1 |
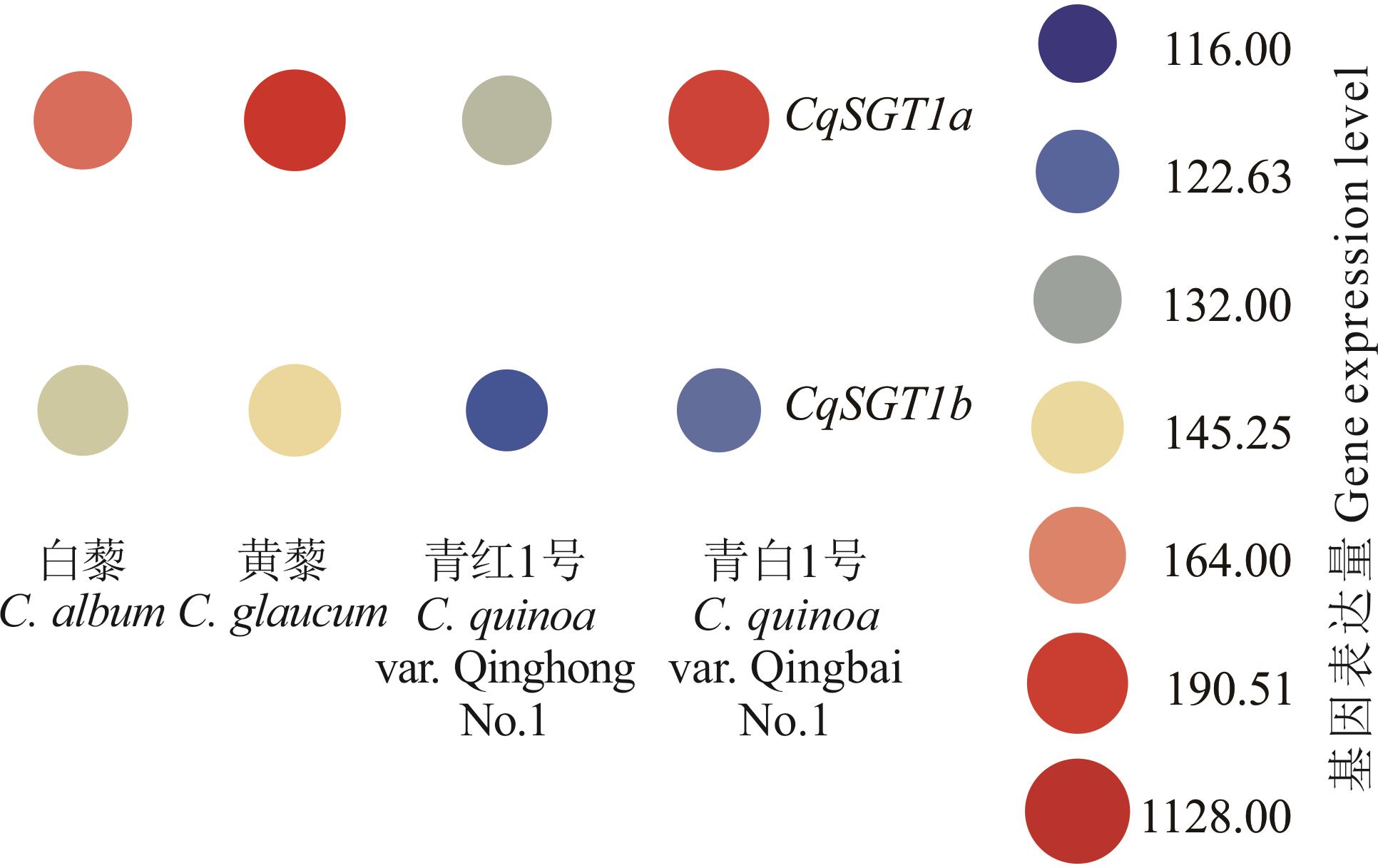
Fig.8 Expression analysis of the CqSGT1 genes in the young seeds of C. album, C. glaucum, C. quinoa var. Qinghong No.1, and C. quinoa var.Qingbai No.1 after flowering
基因 Gene | 染色体位置Chromosome location | Ref | Alt | JQ3 | JQ6 | Qingheili-8 | Qingbaili-9 | 影响Effect(其他突变系Other mutant lines) | 改变的密码子 The changed codon |
|---|---|---|---|---|---|---|---|---|---|
| CqSGT1a | Chr06:34577463 | GA | G | GA, GA | GA, GA | GA, GA | G,G | 内含子Intron | GTa/Gat |
| CqSGT1b | Chr07:83675475 | A | AT | N | A, AT | A, AT | N | 上游Upstream | aAt/aAT |
| CqSGT1b | Chr07:83677523 | A | AC | AC, AC | AC, AC | AC, AC | AC, AC | 内含子Intron | aAc/aAC |
| CqSGT1b | Chr07:83679761 | CA | C | C, C | C, C | C, C | CA, C | 内含子Intron | cCA/cCa |
| CqSGT1b | Chr07:83679830 | TTTGTTGTTGTTG | T | TTTGTTGTTGTTG, TTTGTTGTTGTTG | N | N | TTTGTTGTTGTTG, T | 内含子Intron | cTTTGTTGTTGTTGT/cTt |
| CqSGT1b | Chr07:83689890 | A | ATAGCTGGCACAAACGGTGCCTTGT | A, A | A, ATAGCTGGCACAAACGGTGCCTTGT | A, ATAGCTGGCACAAACGGTGCCTTGT | A, ATAGCTGGCA CAAACGGTGCCTTGT | 下游Downstream | aAt/aATAGCTGGCACAAACGGTGCCTTGTt |
| CqSGT1b | Chr07:83690874 | A | AT | AT, AT | A, A | A, A | N | 下游Downstream | tAt/tAT |
| CqSGT1b | Chr07:83691319 | G | GA | GA, GA | GA, GA | GA, GA | GA, GA | 下游Downstream | acGaaa/acGAaa |
Table 6 DNA variation of CqSGT1s among C. quinoa lines
基因 Gene | 染色体位置Chromosome location | Ref | Alt | JQ3 | JQ6 | Qingheili-8 | Qingbaili-9 | 影响Effect(其他突变系Other mutant lines) | 改变的密码子 The changed codon |
|---|---|---|---|---|---|---|---|---|---|
| CqSGT1a | Chr06:34577463 | GA | G | GA, GA | GA, GA | GA, GA | G,G | 内含子Intron | GTa/Gat |
| CqSGT1b | Chr07:83675475 | A | AT | N | A, AT | A, AT | N | 上游Upstream | aAt/aAT |
| CqSGT1b | Chr07:83677523 | A | AC | AC, AC | AC, AC | AC, AC | AC, AC | 内含子Intron | aAc/aAC |
| CqSGT1b | Chr07:83679761 | CA | C | C, C | C, C | C, C | CA, C | 内含子Intron | cCA/cCa |
| CqSGT1b | Chr07:83679830 | TTTGTTGTTGTTG | T | TTTGTTGTTGTTG, TTTGTTGTTGTTG | N | N | TTTGTTGTTGTTG, T | 内含子Intron | cTTTGTTGTTGTTGT/cTt |
| CqSGT1b | Chr07:83689890 | A | ATAGCTGGCACAAACGGTGCCTTGT | A, A | A, ATAGCTGGCACAAACGGTGCCTTGT | A, ATAGCTGGCACAAACGGTGCCTTGT | A, ATAGCTGGCA CAAACGGTGCCTTGT | 下游Downstream | aAt/aATAGCTGGCACAAACGGTGCCTTGTt |
| CqSGT1b | Chr07:83690874 | A | AT | AT, AT | A, A | A, A | N | 下游Downstream | tAt/tAT |
| CqSGT1b | Chr07:83691319 | G | GA | GA, GA | GA, GA | GA, GA | GA, GA | 下游Downstream | acGaaa/acGAaa |
| [1] | Gómez-Caravaca A M, Iafelice G, Verardo V, et al. Influence of pearling process on phenolic and saponin content in quinoa (Chenopodium quinoa Willd). Food Chemistry, 2014, 15(157): 174-178. |
| [2] | Jiang X D, Li X F, Hao Y P, et al. Gene cloning and express of squalene synthase and β-amyrin synthase from Chenopodium quinoa. Soils, 2018, 50(6): 1214-1221. |
| 姜晓东, 李新凤, 郝艳平, 等. 藜麦β-香树酯醇合酶和鲨烯合酶基因的克隆与表达. 土壤, 2018, 50(6): 1214-1221. | |
| [3] | Jarvis D E, Ho Y S, Lightfoot D J, et al. The genome of Chenopodium quinoa. Nature, 2017, 542(7641): 307-312. |
| [4] | Bazile D, Fuentes F, Mujica A. Historical perspectives and domestication//Bhargava A. Quinoa: botany, production and uses. Wallingford: Centre for Agriculture and Bioscience International, 2013: 16-35. |
| [5] | Hinojosa L, González J A, Barrios-Masias F H, et al. Quinoa abiotic stress responses: a review. Plants, 2018, 7(4): 106. |
| [6] | Wang C. Study on resistance evaluation of quinoa germplasms to downy mildew and its resistance mechanisms. Lanzhou: Gansu Agricultural University, 2023. |
| 王昶. 藜麦种质对霜霉病的抗性评价及其抗病机理研究. 兰州: 甘肃农业大学, 2023. | |
| [7] | Wang C, Li M Q, Yang F R, et al. Diseases investigation and pathogen identification of quinoa downy mildew in Gansu Province. Journal of Nuclear Agriculture, 2023(3): 503-512. |
| 王昶, 李敏权, 杨发荣, 等. 甘肃藜麦霜霉病调查及其病原菌鉴定. 核农学报, 2023(3): 503-512. | |
| [8] | Yuan C L, Li C J, Zhao X B, et al. Genome-wide identification and characterization of HSP90-RAR1-SGT1-Complex members from Arachis genomes and their responses to biotic and abiotic stresses. Frontiers in Genetics, 2021, 12: 689669. |
| [9] | Kitagawa K, Skowyra D, Elledge S J, et al. SGT1 encodes an essential component of the yeast kinetochore assembly pathway and a novel subunit of the SCF ubiquitin ligase complex. Molecular Cell, 1999, 4(1): 21-33. |
| [10] | Wang K, Zhang Z Y. Progress in studying the function of SGT1 in plant disease resistance response. Journal of Plant Genetic Resources, 2008, 9(1): 115-118. |
| 王凯, 张增燕. SGT1在植物抗病反应中的功能研究进展. 植物遗传资源学报, 2008, 9(1): 115-118. | |
| [11] | Zhang D L, Yang X X, Wen Z Y, et al.Proxitome profiling reveals a conserved SGT1-NSL1 signaling module that activates NLR-mediated immunity. Molecular Plant, 2024, 17(9): 1369-1391. |
| [12] | Azevedo C, Sadanandom A, Kitagawa K, et al. The RAR1 interactor SGT1, an essential component of R gene-triggered disease resistance. Science, 2002, 295(5562): 2073-2076. |
| [13] | Wang K, Uppalapati S R, Zhu X H, et al. SGT1 positively regulates the process of plant cell death during both compatible and incompatible plant-pathogen interactions. Molecular Plant Pathology, 2010, 11(5): 597-611. |
| [14] | Makoto I, Ohnishi K, Hikichi Y, et al. Molecular chaperons and co-chaperons, Hsp90, RAR1, and SGT1 negatively regulate bacterial wilt disease caused by Ralstonia solanacearum in Nicotiana benthamiana. Plant Signaling & Behavior, 2015, 10: e970410. |
| [15] | Kumar D, Kirt P B. Pathogen-induced SGT1of Arachis diogoi induces cell death and enhanced disease resistance in tobacco and peanut. Plant Biotechnology Journal, 2015, 13: 73-84. |
| [16] | Yu G, Xian L, Zhuang H Y, et al. SGT1 is not required for plant LRR-RLK-mediated immunity. Molecular Plant Pathology, 2021, 22(1): 145-150. |
| [17] | Chen Z Q, Wu Q, Tong C, et al. Characterization of the roles of SGT1/RAR1, EDS1/NDR1, NPR1, and NRC/ADR1/NRG1 in Sw-5b-mediated resistance to tomato spotted wilt virus. Viruses, 2021, 13(8): 1447. |
| [18] | Shanmugam A, Thamilarasan S K, Park J I, et al. Characterization and abiotic stress-responsive expression analysis of SGT1 genes in Brassica oleracea. Genome, 2016, 59: 243-251. |
| [19] | Berens M L, Berry H M, Mine A, et al. Evolution of hormone signaling networks in plant defense. Annual Review of Phytopathology, 2017, 55(1): 401-425. |
| [20] | Váczy K Z, Otto M, Gomba-Tóth A, et al. Botrytis cinerea causes different plant responses in grape (Vitis vinifera) berries during noble and grey rot: diverse metabolism versus simple defence. Frontiers in Plant Science, 2024, 15: 1433161. |
| [21] | Yu G, Xian L, Xue H, et al. A bacterial effector protein prevents MAPK-mediated phosphorylation of SGT1to suppress plant immunity. PLoS Pathogens, 2020, 16(9): e1008933. |
| [22] | Peart J R, Lu R, Sadanandom A, et al. Ubiquitin ligase-associated protein SGT1 is required for host and nonhost disease resistance in plants. Proceedings of the National Academy of Sciences of the United States of America, 2002, 99(16): 10865-10869. |
| [23] | Zhu X D, Yang L F, Chen Y Y, et al. Biological functional analysis of common buckwheat (Fagopyrum esculentum) FeSGT1 gene in enhancing drought stress resistance. Acta Agronomica Sinica, 2023, 49(6): 1573-1583. |
| 朱旭东, 杨兰锋, 陈媛媛, 等. 甜荞FeSGT1基因克隆及抗旱功能解析. 作物学报, 2023, 49(6): 1573-1583. | |
| [24] | Agarwal G, Garg V, Kudapa H, et al. Genome-wide dissection of AP2/ERF and HSP90 gene families in five legumes and expression profiles in chickpea and pigeonpea. Plant Biotechnology Journal, 2016, 14: 1563-1577. |
| [25] | Zou C S, Chen A J, Xiao L H, et al. A high-quality genome assembly of quinoa provides insights into the molecular basis of salt bladder-based salinity tolerance and the exceptional nutritional value. Cell Research, 2017, 27(11): 1327-1340. |
| [26] | Xie Y J, Xue J, Jiang X D, et al. Screening of reference genes in Chenopodium quinoa under Peronospora variabilis stress and verification of their stability. Journal of Fujian Agriculture and Forestry University (Natural Science Edition), 2024, 53(2): 191-198. |
| 解宇洁, 薛婧, 姜晓东, 等. 霜霉病菌胁迫下藜麦内参基因的筛选及其稳定性验证. 福建农林大学(自然科学版), 2024, 53(2): 191-198. | |
| [27] | Koua A P, Oyiga B C, Baig M M, et al. Breeding driven enrichment of genetic variation for key yield components and grain starch content under drought stress in winter wheat. Frontiers in Plant Science, 2021, 12: 684205. |
| [28] | Meldau S, Baldwin I T, Wu J Q. For security and stability: SGT1 in plant defense and development. Plant Signaling & Behavior, 2011, 6(10): 1479-1482. |
| [29] | Holt B F, Belkhadir Y, Dangl J L. Antagonistic control of disease resistance protein stability in the plant immune system. Science, 2005, 309(5736): 929-932. |
| [30] | Muskett P, Parker J. Role of SGT1 in the regulation of plant R gene signaling. Microbes and Infection, 2003, 5(11): 969-976. |
| [31] | Chen X Y, Li X B, Duan Y H, et al. A secreted fungal subtilase interferes with rice immunity via degradation of suppressor of G2 allele of skp1. Plant Physiology, 2022, 190(2): 1474-1489. |
| [32] | Wang Y Q, Liu C, Du Y Y, et al. A stripe rust fungal effector PstSIE1 targets TaSGT1 to facilitate pathogen infection. The Plant Journal, 2022, 112(6): 1413-1428. |
| [33] | Steven S R, Huang L, Brandt A S, et al. Development of a virus-induced gene-silencing system for hexaploid wheat and its use in functional analysis of the Lr21-mediated leaf rust resistance pathway. Plant Physiology, 2005, 138(4): 2165-2173. |
| [34] | Li X B. Functional characterization of subtilisin-like protease family genes in Ustilaginoidea virens. Changsha: Central China Agricultural University, 2022. |
| 李夏冰. 稻曲菌枯草杆菌蛋白酶家族基因功能研究. 长沙: 华中农业大学, 2022. | |
| [35] | Ouyang X, Chen J L, Sun Z M, et al. Ubiquitin E3 ligase activity of Ralstonia solanacearum effector RipAW is not essential for induction of plant defense in Nicotiana benthamiana. Frontiers in Microbiology, 2023, 14: 1201444. |
| [36] | Zhang C L, Xu D C, Jiang X C. Improvement of disease resistance of sugar beet by molecular breeding. China Beet & Sugar, 2008(2): 23-26. |
| 张春来, 徐德昌, 姜孝成. 分子育种提高甜菜抗病性. 中国甜菜糖业, 2008(2): 23-26. | |
| [37] | Guo W L, Chen B H, Guo Y Y, et al. Improved powdery mildew resistance of transgenic Nicotiana benthamiana overexpressing the Cucurbita moschata CmSGT1 gene. Frontiers in Plant Science, 2019, 10: 955. |
| [38] | Forner-Giner M Á, Rodríguez-Gamir J, Primo-Millo E, et al. Hydraulic and chemical responses of citrus seedlings to drought and osmotic stress. Journal of Plant Growth Regulation, 2011, 30(3): 353-366. |
| [39] | Luna C M, Pastori G M, Driscoll S, et al. Drought controls on H2O2 accumulation, catalase (CAT) activity and CAT gene expression in wheat. Journal of Experimental Botany, 2005, 56(411): 417-423. |
| [40] | Baczek-Kwinta R, Filek W, Grzesiak S, et al. The effect of soil drought and rehydration on growth and antioxidative activity in flag leaves of triticale. Biologia Plantarum, 2006, 50(1): 55-60. |
| [41] | Mhamdi A, Queval G, Chaouch S, et al. Catalase function in plants: a focus on Arabidopsis mutants as stress-mimic models. Journal of Experimental Botany, 2010, 61(15): 4197-4220. |
| [42] | Jiang J M, Chen J T, Luo L, et al. Expression analysis reveals that sorghum disease resistance protein SbSGT1 is regulated by auxin. Biology, 2022, 11(1): 67. |
| [43] | Li R Q, Zheng W Y, Yang R F, et al. OsSGT1promotes melatonin-ameliorated seed tolerance to chromium stress by affecting the OsABI5-OsAPX1 transcriptional module in rice. The Plant Journal, 2022, 112(1): 151-171. |
| [44] | Noël L D, Cagna G, Stuttmann J, et al. Interaction between SGT1 and cytosolic/nuclear HSC70 chaperons regulates Arabidopsis immune responses. Plant Cell, 2007, 19(12): 4061-4076. |
| [1] | Wei-peng ZOU, Yi LIU, Jia-xing ZHAI, Si-yi ZHOU, Zhi-yi GONG, Hui-fang CEN, Hui-sen ZHU, Tao XU. Cloning of MsNAC053 from alfalfa and analysis of its transcript profile in response to abiotic stresses [J]. Acta Prataculturae Sinica, 2025, 34(9): 121-133. |
| [2] | Chang WANG, Geng-mei MIN, Li-juan ZHANG, Jian-ying LU, Zao-xia NIU, Yu-ming WEI, Fa-rong YANG. Progress in research on quinoa downy mildew and its integrated control [J]. Acta Prataculturae Sinica, 2025, 34(9): 194-205. |
| [3] | Yuan-yuan ZHAO, Xiao-jian PU, Cheng-ti XU, Wei WANG, Yun-jie FU. Cloning of the MtBMI1 gene from Medicago truncatula and its role in drought tolerance [J]. Acta Prataculturae Sinica, 2025, 34(6): 139-153. |
| [4] | Xin-yue ZHOU, Li-ping WANG, Qing-xue JIANG, Xiao-ran MA, Deng-xia YI, Xue-min WANG. Isolation of the low-temperature induced proteinMsLTI65 from alfalfa and its response to different stresses [J]. Acta Prataculturae Sinica, 2025, 34(5): 89-104. |
| [5] | Xin-yao WANG, Ya-ping PENG, Li-rong YAO, Jun-cheng WANG, Er-jing SI, Hong ZHANG, Ke YANG, Xiao-le MA, Ya-xiong MENG, Hua-jun WANG, Bao-chun LI. Gene cloning and drought resistance identification of the gene HgS5 in Halogeton glomeratus [J]. Acta Prataculturae Sinica, 2025, 34(2): 184-195. |
| [6] | Yong-long LI, Sheng-hui ZHOU, Meng-yao XUE, Yuan GAO, Le JU, Yi-bing CHEN, Song-lin FU, Jian-hao HAO, Heng LI, Kun ZHANG, Zhi-fang ZUO. Cloning of the gene ZjWRKY63 from Zoysia japonica and its salt resistance analysis in transgenic Arabidopsis [J]. Acta Prataculturae Sinica, 2025, 34(12): 157-169. |
| [7] | Lin BIAN, Yan ZHANG, Xiao-wei HUO, Rui DAI, Na GUO, Feng-yan YI, Cui-ping GAO, Zhi-qiang ZHANG. Identification of the CKX gene family in alfalfa and its responses to abiotic stress [J]. Acta Prataculturae Sinica, 2025, 34(11): 125-135. |
| [8] | Long-yi HE, Meng-meng TAN, Hai-tao CHE, Hong-ying ZHANG, Yu-xin ZHU, Yan-ni ZHANG. Cloning and analysis of drought tolerance function of the LpDREB9 in Lilium pumilum [J]. Acta Prataculturae Sinica, 2025, 34(1): 161-173. |
| [9] | Yi WU, Ya-lan FENG, Tian-ning WANG, Ji-hao JU, Hui-shu XIAO, Chao MA, Jun ZHANG. Genome-wide identification and expression analysis of the Hsp70 gene family in wheat and its ancestral species [J]. Acta Prataculturae Sinica, 2024, 33(7): 53-67. |
| [10] | Zhen-huan ZHANG, Li-rong YAO, Jun-cheng WANG, Er-jing SI, Hong ZHANG, Ke YANG, Xiao-le MA, Ya-xiong MENG, Hua-jun WANG, Bao-chun LI. Identification of AKR gene family members in Halogeton glomeratus and salt tolerance analysis of the root salt stress response gene HgAKR42639 [J]. Acta Prataculturae Sinica, 2024, 33(7): 68-83. |
| [11] | Hai-ming KONG, Jia-xing SONG, Jing YANG, Qian LI, Pei-zhi YANG, Yu-man CAO. Identification and transcript profiling of the CAMTA gene family under abiotic stress in alfalfa [J]. Acta Prataculturae Sinica, 2024, 33(5): 143-154. |
| [12] | Ze-bin LI, Yong-zheng QIU, Yan-jie LIU, Jin-qiu YU, Bai-ji WANG, Qian-ning LIU, Yue WANG, Guo-wen CUI. Identification of the BZR gene family in alfalfa and analysis of its transcriptional responses to abiotic stress [J]. Acta Prataculturae Sinica, 2024, 33(11): 106-122. |
| [13] | Xin-yue ZHOU, Qing-xue JIANG, Hui-li JIA, Lin MA, Lu FAN, Xue-min WANG. Cloning and salt-tolerance functional analysis of alfalfa MsBBX20 gene [J]. Acta Prataculturae Sinica, 2024, 33(10): 55-73. |
| [14] | Shang-qin HU, Jun-cheng WANG, Li-rong YAO, Er-jing SI, Xiao-le MA, Ke YANG, Hong ZHANG, Ya-xiong MENG, Hua-jun WANG, Bao-chun LI. Cloning and preliminary functional analysis of the root gene HgAKR6C of Halogeton glomeratus [J]. Acta Prataculturae Sinica, 2024, 33(1): 61-74. |
| [15] | Zhen-fen ZHANG, Rong HUANG, Xiang-yang LI, Bo YAO, Gui-qin ZHAO. Seed-borne bacterial diversity of oat and functional analysis based on Illumina MiSeq high-throughput sequencing [J]. Acta Prataculturae Sinica, 2023, 32(7): 96-108. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||